To see some examples of the natverse in action take a look at our Gallery. Alternatively if you’d rather see some code right away, then read one of the introductory articles. If these look interesting, then we recommend reading (or at least skimming) the natverse paper for a more in depth introduction. You can then browse documentation for the core nat package and work through some example code.
natverse paper
To understand what the natverse can do and how it is organised, the best resource is a paper currently available at elife : Bates, Manton et al (2020). This describes all the main natverse packages and how they fit together. It also provides numerous worked examples based on data from a range of species, including flies, fish and mice.
nat documentation
To get started with some actual code, take a look at the documentation for the nat (NeuroAnatomy Toolbox) package, which is the core of the natverse:
- Read the overview package documentation
(
?natin R) - Read the Introduction to neurons article
- There are several other useful articles.
- Check out the thematically organised function reference documentation.
- Most help pages include examples.
There is extensive sample code and data available including:
- nat.examples has detailed examples for data sets from a range of model organisms and techniques
- frulhns analysis of sexually dimorphic circuits
Bridging and Mirroring registrations
We introduced the concept of bridging registrations between template brains in Manton et al, bioRxiv, 2014. This allows a huge amount of neuroanatomical data from different labs acquired using both light and electron microscopy to brought into a common space for shared analysis. The natverse provides efficient tools to move data from one template brain to another. The original version of this paper introduces and motivates our general approach with many examples. The natverse supports a range of different registration tools and formats including thin plate splines, CMTK, ANTs and the Saalfeld lab’s h5 format.
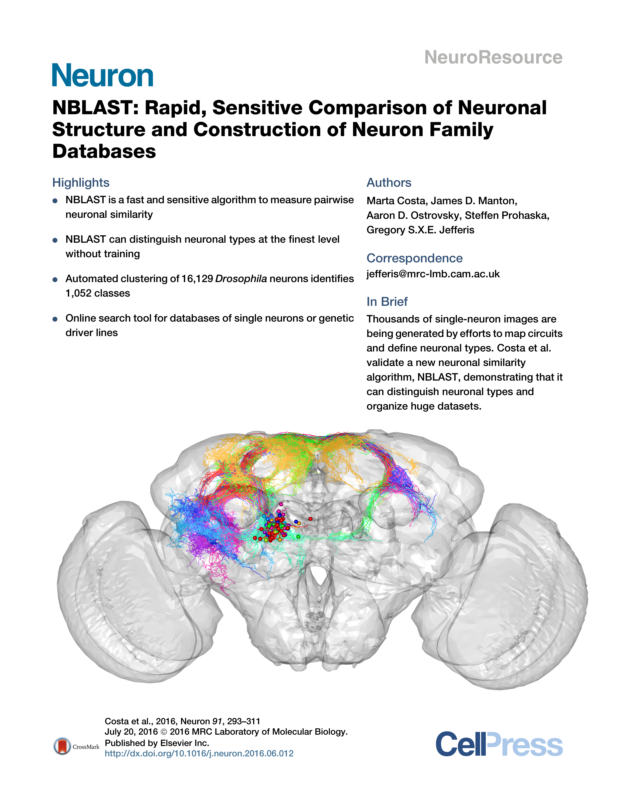
NBLAST
NBLAST is a fast and sensitive algorithm for comparing neuron structure. It is described in an open access publication, Costa et al, Neuron (2016). Read the paper to understand how the algorithm works and to see many examples of its use.
You can also see the R Markdown reports used to generate all the figures for the paper.
Learning R
Naturally you will need to learn some basics of the R language to use the natverse. There are a huge number of books and online resources now available. The tidyverse website has some good suggestions, see https://www.tidyverse.org/learn/. Resources that we recommend include:
- http://swirlstats.com (an interactive way to get started for complete beginners)
- https://www.rstudio.com/resources/cheatsheets/ (a quick way to find key functions for data munging tasks)
- http://adv-r.had.co.nz (chapters 1-7 are a good investment if you plan to spend much time in R)
By: