prune_vertices removes vertices from a neuron
prune_edges removes edges (and any unreferenced vertices)
prune_vertices(x, verticestoprune, invert = FALSE, ...)
prune_edges(x, edges, invert = FALSE, ...)Arguments
- x
A
neuronto prune. This can be any object that can be converted byas.ngraph--- see details.- verticestoprune
An integer vector describing which vertices to remove.
- invert
Whether to keep vertices rather than dropping them (default FALSE).
- ...
Additional arguments passed to
as.neuron.ngraph- edges
The edges to remove. One of i) an Nx2 matrix, each row specifying a single edge defined by its raw edge id, ii) an integer vector defining a path of raw vertex ids or iii) an
igraph.esedge sequence --- see details and thePandpatharguments ofigraph::E.
Value
A pruned neuron
Details
These are relatively low-level functions and you will probably want
to use subset.neuron or prune.neuron and
friends in many cases.
Note that prune_vertices and prune_edges both use raw
vertex ids to specify the vertices/edges to be removed. If you want to use
the id in the PointNo field, then you must translate yourself (see
examples).
Both prune_vertices and prune_edges first convert their input
x to the ngraph representation of the neuron before
removing points. The input x can therefore be in any form compatible
with as.ngraph including an existing ngraph. There is
an additional requirement that the input must be compatible with
xyzmatrix if invert=TRUE.
Note that the edges argument of prune_edges must specify a
path traversing a set of vertices in a valid order. However if the input is
a matrix or vector the direction of each individual edge in this path is
ignored. So if your neuron has edges 2->1 2->3 3->4 then an edge sequence
1:3 would successfully delete 2 edges.
See also
Examples
n=prune_vertices(Cell07PNs[[1]], 1:25)
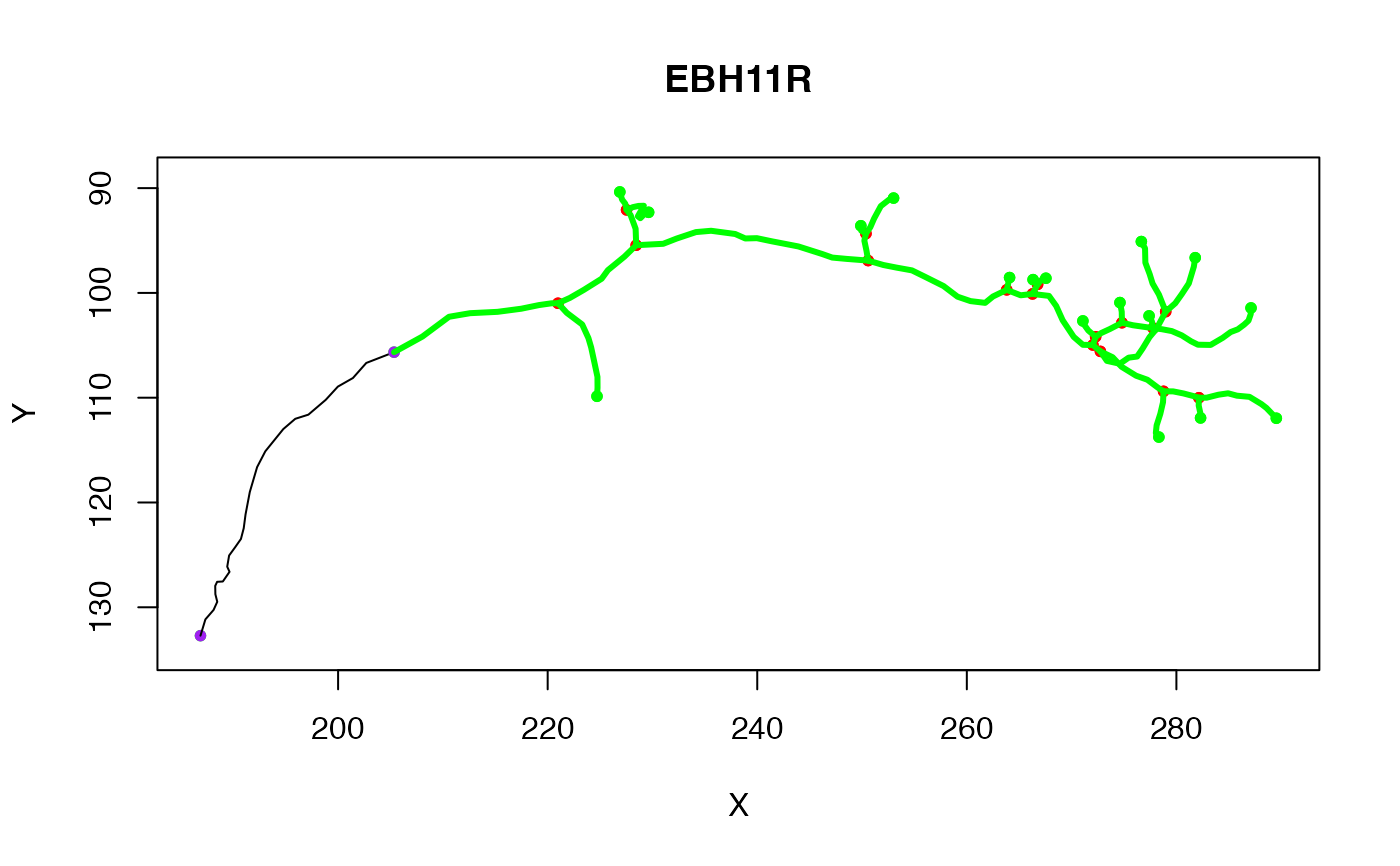
# original neuron
plot(Cell07PNs[[1]])
# with pruned neuron superimposed
plot(n, col='green', lwd=3, add=TRUE)
 # use the PointNo field (= the original id from an SWC file)
n2=prune_vertices(n, match(26:30, n$d$PointNo))
y=prune_edges(Cell07PNs[[1]], edges=1:25)
# remove the spine of a neuron
spine_ids=spine(Cell07PNs[[1]], rval='ids')
pruned=prune_edges(Cell07PNs[[1]], spine_ids)
# NB this is subtly different from this, which removes vertices along the
# spine *even* if they are part of an edge that is outside the spine.
pruned2=prune_vertices(Cell07PNs[[1]], spine_ids)
# use the PointNo field (= the original id from an SWC file)
n2=prune_vertices(n, match(26:30, n$d$PointNo))
y=prune_edges(Cell07PNs[[1]], edges=1:25)
# remove the spine of a neuron
spine_ids=spine(Cell07PNs[[1]], rval='ids')
pruned=prune_edges(Cell07PNs[[1]], spine_ids)
# NB this is subtly different from this, which removes vertices along the
# spine *even* if they are part of an edge that is outside the spine.
pruned2=prune_vertices(Cell07PNs[[1]], spine_ids)