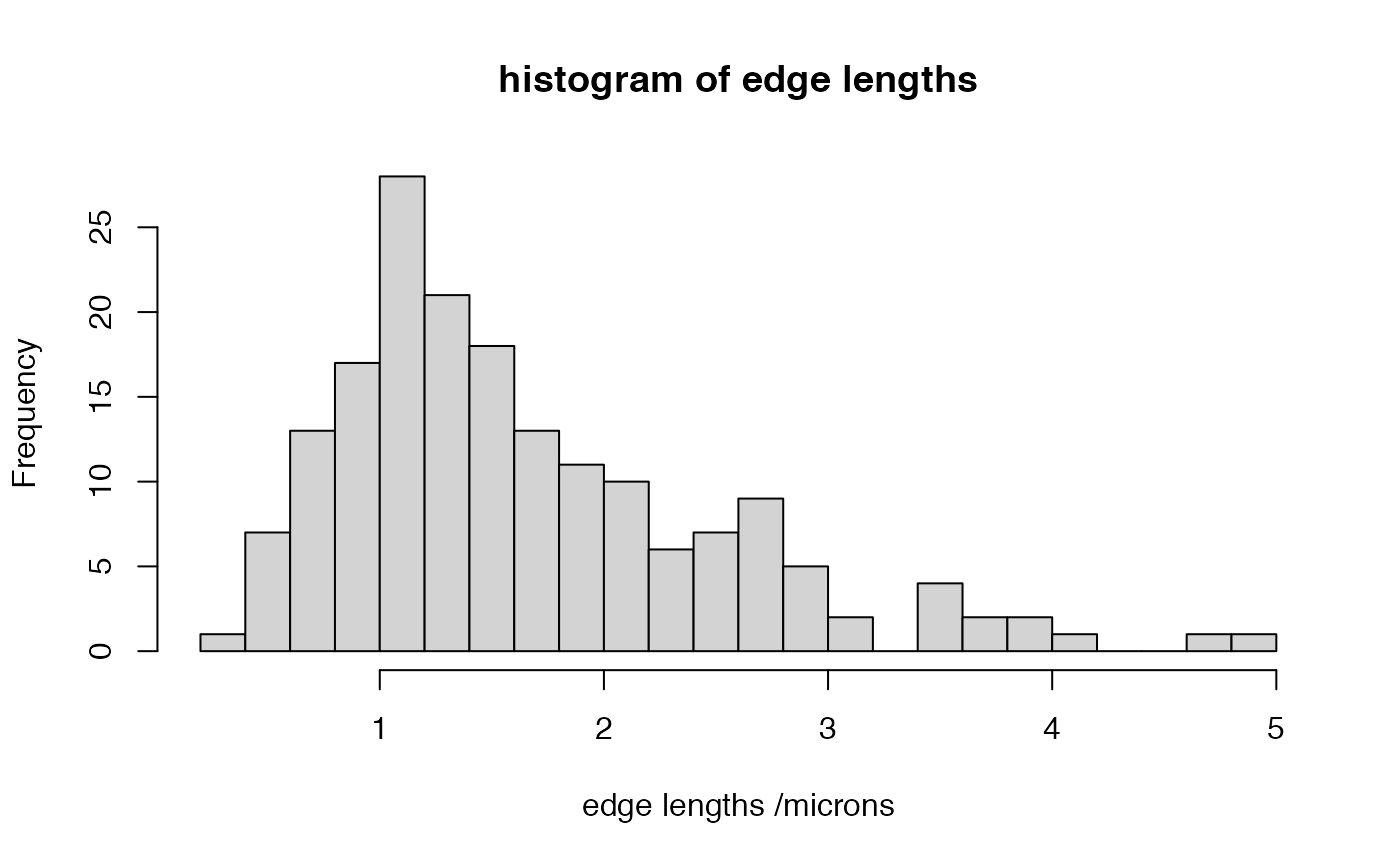
Calculate length of all segments in neuron
seglengths(x, all = FALSE, flatten = TRUE, sumsegment = TRUE)Arguments
- x
A neuron
- all
Whether to calculate lengths for all segments when there are multiple subtrees (default:
FALSE)- flatten
Whether to flatten the lists of lists into a single list when
all=TRUE- sumsegment
Whether to return the length of each segment (when sumsegment=TRUE, the default) or a list of vectors of lengths of each individual edge in the segment.
Value
A vector of lengths for each segment or when
sumsegment=FALSE a list of vectors
Details
A segment is an unbranched portion of neurite consisting of at least
one vertex joined by edges.Only segments in x$SegList will be calculated
unless all=TRUE. Segments containing only one point will have 0
length.