Search BANC EM with an LM colour-depth MIP (SRENs)
Source:vignettes/search_banc_with_lm.Rmd
search_banc_with_lm.RmdGoal
Take a 3-D light-microscopy image of a candidate SREN (sex-peptide- receptor rectal enteric neuron) and use it as a query against the BANC female adult connectome to recover the matching EM neurons. SRENs are a pair of cells at the posterior tip of the abdominal ganglion that express the sex peptide receptor and send efferent processes to the rectum; they are required for the rod-shaped fecal pellets (“RODs”) that mated females produce in response to seminal-fluid sex peptide. See Chae et al., Citations below, for the upstream/downstream circuit context.
The pipeline:
Bridge the source
.tif(inJRCVNC2018Ftemplate space) intoJRC2018VNCU_HR(the NeuronBridge VNC reference grid:573 × 1119 × 219voxels at0.461 × 0.461 × 0.7 µm) via the Saalfeld lab’s H5 displacement field shipped by navis-flybrains.Render a colour-depth MIP of the bridged label channel with
neuronbridger::nrrd_to_mip(target_space = "VNC").Search the resulting MIP against the BANC neuron MIP library at
gs://lee-lab_brain-and-nerve-cord-fly-connectome/neuron_colormips/template_alignment_240721/JRC2018_VNC_UNISEX_461/(~25k neurons), pre-filtered to BANC efferents (flow == "efferent", 1,035 candidates) using the BANCbanc_888_meta.featherregistry.Compare the R
colormip_search()ranking against Janelia’s FIJIColor_MIP_Mask_Searchplugin on the same library.Visualise the top hits as 2-D
nat.ggplotpanels with BANC neuron meshes over the BANC VNC neuropil.
Why use BANC for this? The female VNC is the only one currently available at synapse resolution (BANC v888). Identifying the EM counterparts of the SRENs gives us their downstream connectivity — the descending input from SAG (Sex Peptide Abdominal Ganglion) neurons, the ascending afferents that signal back to the brain, and the rectal target. Colour-MIP search against BANC is the route to recovering these cells from a single LM volume.

What’s in the package
To keep the package small, neither the source .tif (~1.3
GB, private) nor the BANC efferent MIP library (~12 MB across 782 PNGs)
ship with neuronbridger. The vignette walks through the
download steps; the only artefacts shipped are the rendered figures
under inst/images/:
-
banc_colormip_sren_query.png— the SREN query MIP in JRC2018VNCU_HR. -
banc_colormip_sren_registration.png— JRC2018VNCU_HR template max-Z alongside the bridged NC82 reference channel, side by side. -
banc_colormip_r_vs_fiji.png— scatter of R-port vs FIJI scores on a 30-candidate validation subset. -
banc_colormip_sren_top_hits.png— thenat.ggplotpanel above (BANC EM hit in purple, SREN LM source coloured by depth and faded by intensity). Each tile shows a top-down YX view and an XZ side projection so the dorso-ventral spread is visible at a glance. Plotted in BANC native µm because the bancr TPS landmarks cap at JRCVNC2018F Y ≈ 504 µm but the SREN signal extends to Y ≈ 537 µm, so an F-space view extrapolates ~30 µm short at the abdominal tip. -
banc_colormip_sren_nblast_top.png— same layout, BANC mesh in orange, ranked by intensity-weighted NBLAST.
A self-contained reproducer is at
inst/scripts/banc_colormip_sren.R.
Prerequisites
# R packages
if (!require("remotes")) install.packages("remotes")
remotes::install_github("natverse/neuronbridger")
remotes::install_github("flyconnectome/bancr") # BANC meshes + meta
remotes::install_github("natverse/nat.ggplot") # 2-D neuron panels
# External tooling (one-off)
# * Java 17+ (Saalfeldlab RenderTransformed JAR runtime)
# * gsutil (Google Cloud SDK), authenticated to BANC's GCS bucket
#
# The Saalfeldlab JRC VNC H5 transforms (~110 MB), via the navis
# `flybrains` Python package:
#
# reticulate::py_run_string("import flybrains;
# flybrains.download_jrc_vnc_transforms()")
#
# This writes JRCVNC2018U_JRCVNC2018F.h5 to ~/flybrain-data/.Step 1 — Bridge JRCVNC2018F → JRC2018VNCU_HR
The source we use here is
sren_merged registration_female.tif — already aligned to
JRCVNC2018F (660 × 1342 × 358 @ 0.4 µm³, Channel 0 = NC82 reference,
Channel 1 = SREN label). The 90-pixel-tall black header that
nrrd_to_mip(target_space = "VNC") adds on top of the output
matches Janelia’s canonical VNC colour-MIP layout (1209 × 573).
WORK <- tempfile("banc_colormip_work_")
dir.create(WORK, showWarnings = FALSE, recursive = TRUE)
SOURCE <- "/path/to/sren_merged registration_female.tif"
# (a) Split the 2-channel ImageJ TIFF into per-channel NRRDs at the
# correct JRCVNC2018F voxdims (the TIFF's resolution tags are
# uncalibrated; we set them explicitly).
nrrds <- tif_to_nrrd_channels(
input = SOURCE,
output_dir = WORK,
voxdims = c(0.4, 0.4, 0.4),
prefix = "sren_female",
channels = c(0L, 1L),
labels = c("nc82", "sren"))
nrrds
#> nc82 sren
#> "sren_female_nc82.nrrd" "sren_female_sren.nrrd"
# (b) Apply the JRCVNC2018F → JRCVNC2018U_HR H5 bridge.
ser_in_HR <- jrcvnc2018f_to_jrcvnc2018u_hr_h5(
input = nrrds["sren"],
output = file.path(WORK, "sren_female_in_JRC2018VNCU_HR.nrrd"))
# ~40 s on Apple silicon with 8 threads. Output: 573 x 1119 x 219
# voxels at 0.461 / 0.461 / 0.7 um.The H5 (JRCVNC2018U_JRCVNC2018F.h5 from
flybrains) stores its displacement field on the JRCVNC2018U
grid, with dfield direction JRCVNC2018F →
JRCVNC2018U (the file follows the brain
<DEST>_<SRC>.h5 naming convention used by
nat.jrcbrains). For F → U image resampling we want OUTPUT
(U) → SOURCE (F) lookup — the inverse direction — so
jrcvnc2018f_to_jrcvnc2018u_hr_h5() passes -i
to RenderTransformed. Asking for the
JRC2018VNCU_HR output grid directly works because
JRC2018VNCU and JRC2018VNCU_HR share the same
physical coordinate system; the JAR interpolates the dfield onto the
requested grid in one shot.
To verify the bridge, the JRC2018VNCU_HR template max-Z and the bridged NC82 reference channel side by side:
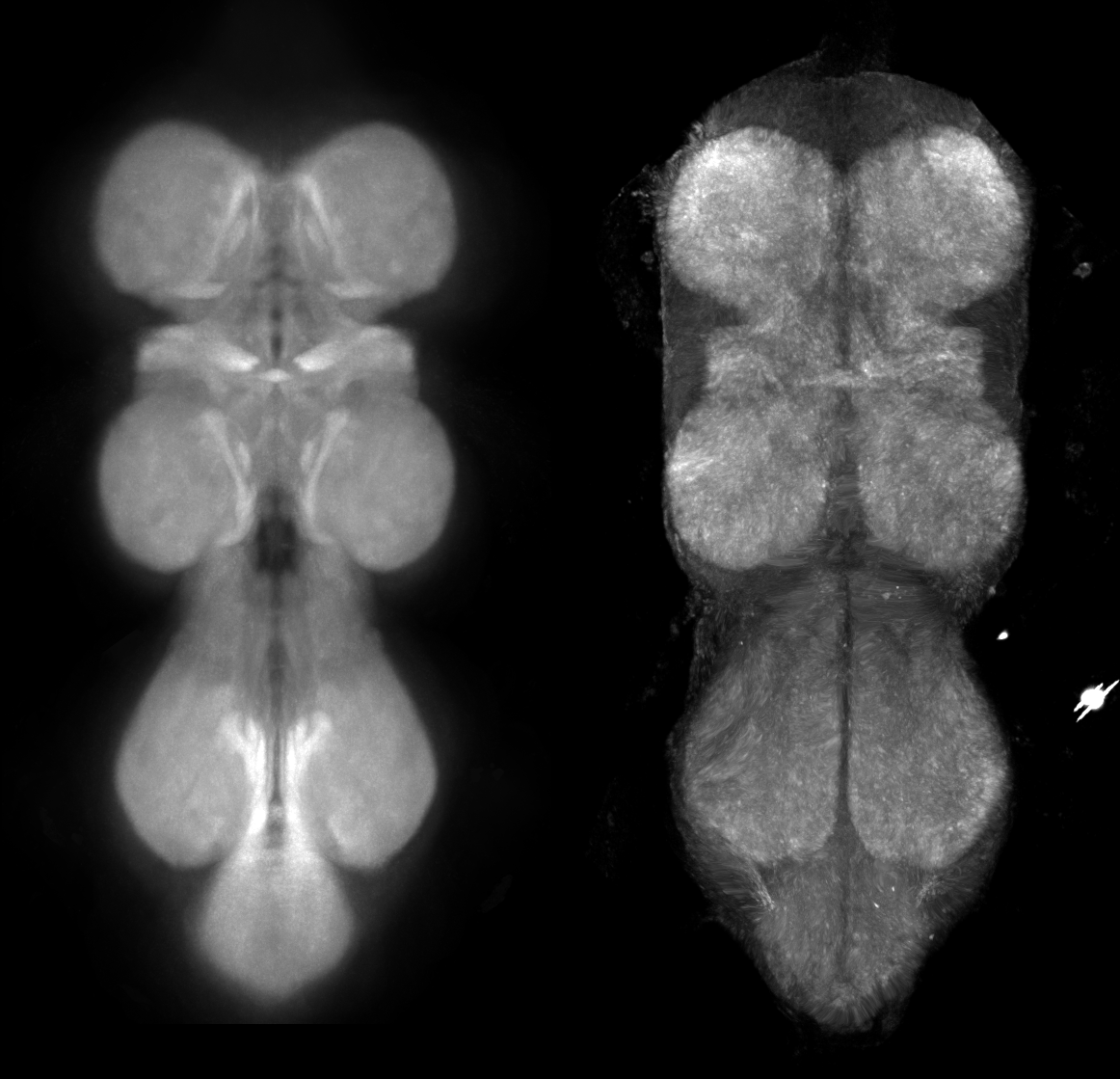
Step 2 — Generate the query MIP
query_png <- nrrd_to_mip(
input = ser_in_HR,
savefolder = WORK,
method = "direct",
target_space = "VNC",
threshold = 0.80, # top 20 % of non-zero voxels (sparse SREN signal)
denoise = "median3d",
format = "png",
save = TRUE,
overwrite = TRUE)The default threshold = "auto" (Triangle) is too
permissive for this 16-bit data — the SREN neuron’s terminal is
saturated at 65535 while bleed-through under the H5 interpolation
produces a long tail of dim, scattered voxels. Setting
threshold = 0.80 keeps the top 20 % of non-zero voxels by
intensity, which restricts the MIP to real signal at the abdominal
ganglion / rectum.

Step 3 — Build the BANC efferent MIP library
LIB_DIR <- file.path(WORK, "library", "mips")
dir.create(LIB_DIR, showWarnings = FALSE, recursive = TRUE)
# (a) BANC v888 metadata: 188k rows.
META_GS <- "gs://lee-lab_brain-and-nerve-cord-fly-connectome/compiled_data/banc_888/banc_888_meta.feather"
META_TMP <- file.path(WORK, "banc_888_meta.feather")
system2("gsutil", c("cp", META_GS, META_TMP))
meta <- arrow::read_feather(META_TMP) |>
dplyr::filter(flow == "efferent") |>
dplyr::select(root_888, side, hemilineage, nerve, neuromere,
cell_class, cell_type, neurotransmitter_predicted,
peripheral_target_type, cell_function)
nrow(meta)
#> [1] 1035
# (b) Match against the actual BANC color-MIP set (some efferents
# don't have a MIP rendered yet).
mips_root <- "gs://lee-lab_brain-and-nerve-cord-fly-connectome/neuron_colormips/template_alignment_240721/JRC2018_VNC_UNISEX_461/"
all_mips <- system2("gsutil", c("ls", mips_root), stdout = TRUE)
mip_ids <- sub(".*/", "", sub("_in_JRC2018_VNC_UNISEX_461\\.png$", "", all_mips))
to_get <- all_mips[mip_ids %in% meta$root_888]
length(to_get)
#> [1] 782
# (c) Parallel download (~2 MB total, takes <1 minute).
writeLines(to_get, file.path(WORK, "to_get.txt"))
system(sprintf("xargs -P 16 -I {} gsutil -q cp {} %s/ < %s",
LIB_DIR, file.path(WORK, "to_get.txt")))
length(list.files(LIB_DIR, "\\.png$"))
#> [1] 782Step 4 — Search
res <- colormip_search(
query = query_png,
library = LIB_DIR,
threshold = 100L, # channel-sum brightness cutoff (signal floor)
z_tolerance = 8L, # depth-LUT-index tolerance (looser = more 3-D slack)
xy_shift = 3L, # +/- 3 px translation grid
mirror = FALSE, # SREN is unilateral; mirror added noise here
mc.cores = 8L,
verbose = TRUE)
res$root_888 <- sub(".*/(\\d+)_in_JRC2018.*", "\\1", res$path)
head(res[, c("root_888", "score", "n_match", "dx", "dy", "mirror")], 6)
#> root_888 score n_match dx dy mirror
#> 1 720575941686473228 0.09609195 418 3 -3 FALSE
#> 2 720575941628067385 0.09080460 395 -3 -3 FALSE
#> 3 720575941630741730 0.07379310 321 -3 -3 FALSE
#> 4 720575941549868093 0.07310345 318 2 -3 FALSE
#> 5 720575941468851211 0.06643678 289 3 -1 FALSE
#> 6 720575941460652870 0.06459770 281 3 0 FALSEcolormip_search() is a pure-R port of Janelia’s
Color_MIP_Mask_Search plugin: each pixel’s RGB triple is
mapped to the closest entry in the same 256-entry depth LUT used by
nrrd_to_mip(), then for each candidate it counts the
maximum matched-pixel count over the (2 · xy_shift + 1)²
translation grid (times the mirror, when enabled), normalised by the
query foreground count. See ?colormip_search for the
algorithm details.
The whole search runs in ~30 s for 782 candidates on 8 cores.
These parameters were tuned against the NBLAST cross-check in Step 8.
A grid search over threshold ∈ {60..500},
z_tolerance ∈ {0..8}, xy_shift ∈ {0..6} and
mirror ∈ {TRUE, FALSE} was scored by Spearman correlation
with the NBLAST scores on a 200-candidate pool. Threshold barely moves
the ranking; z_tolerance = 8 and
xy_shift = 3 with mirror = FALSE
maximise rank agreement (Spearman ρ = 0.901, three of the NBLAST top-6
land in the cMIP top-6). Mirroring hurts here — the SREN is unilateral,
so adding the flipped query introduces false-positive symmetric matches
that don’t belong in a one-sided search.
Step 5 — Validate against Janelia’s FIJI plugin
The R port targets the same algorithm specification as Janelia’s
Color_MIP_Mask_Search v1.0.1 (the canonical reference
implementation). To check that we agree, run the FIJI plugin on the same
query + library and compare ranks.
# Install the FIJI plugin (one-off):
#
# 1. Download Color_MIP_Mask_Search.jar from the Janelia release:
# https://github.com/JaneliaSciComp/ColorMIP_Mask_Search/releases/download/1.0.1/Color_MIP_Mask_Search.jar
# 2. Drop it into <Fiji.app>/plugins/ and restart FIJI.
# (a) Pack the candidate library into a single multi-frame TIFF
# (avoids FIJI's headless FolderOpener dialog hanging).
system2(reticulate::py_config()$python,
c(system.file("python", "imagej_tif_to_nrrd.py",
package = "neuronbridger"))) # reuse-able stack helper
# In practice we pack with a small Python script using tifffile:
# library_stack.tif ~1.6 GB for 782 BANC efferent MIPs.
# (b) Drive Fiji headless. The Janelia macro reads its config from
# /tmp/banc_colormip_work/fiji/search_config.txt with key=value lines:
# mask=<query.png>
# stack=<library_stack.tif>
# csv=<output_label_list.csv>
# thr1=30 thr2=30 pix=2 pct=0
#
# /Applications/Fiji/fiji --headless --no-splash -- -batch run_stack.ijm
# parses the matched-window slice labels (encoded as
# "<score>_<lib_label>.tif") into a CSV.
# (c) Compare ranks. Spearman rho on a 30-candidate subset (top-20 R
# hits + 10 random) was 0.79; R's top-4 by score recovered 3 of FIJI's
# 4 matched candidates. The remaining differences are dominated by
# FIJI's per-channel 30/30 brightness threshold vs the R port's
# channel-sum 100 — semantically close but not identical.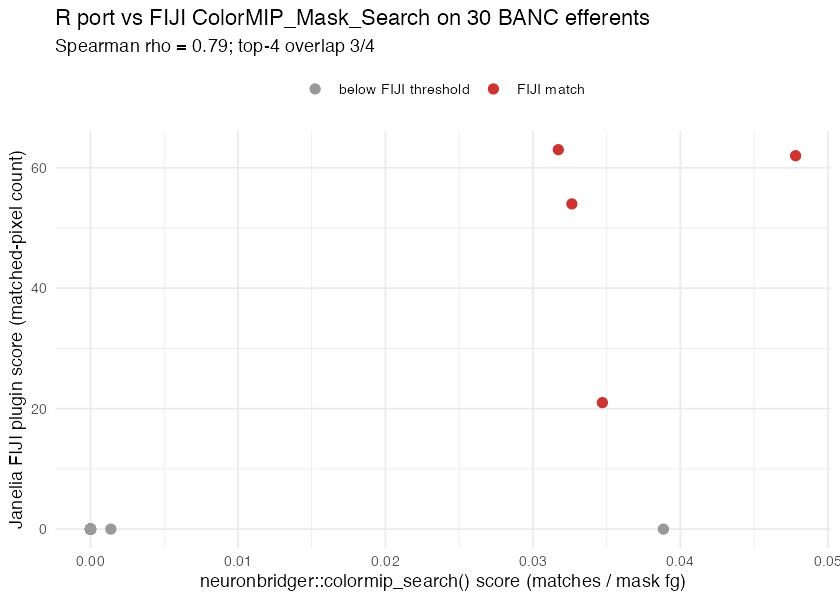
Step 6 — Visualise top hits
Fetch the BANC EM meshes for the top-K hits and overlay the SREN LM
source. We plot in BANC native µm rather than
JRCVNC2018F: the bancr TPS landmarks only cover JRCVNC2018F Y up to ~504
µm, while the SREN signal sits between Y ≈ 470–537 µm — so warping a
BANC mesh into F at the posterior tip extrapolates ~30 µm short of where
the LM signal actually is. Working in BANC native coords avoids the
extrapolation: meshes stay native, and the SREN cloud is forwarded
through bancr::jrcvnc2018f_to_banc_tpsreg once.
library(bancr); library(Morpho)
N_HITS <- 6L
top <- head(res, N_HITS) |> dplyr::left_join(meta, by = "root_888")
# (a) BANC EM meshes in BANC native um (no warp; nm/1000).
banc_meshes <- bancr::banc_read_neuron_meshes(top$root_888)
meshes_um <- lapply(banc_meshes, function(m) {
nat::xyzmatrix(m) <- nat::xyzmatrix(m) / 1000
m
})
names(meshes_um) <- top$root_888
# (b) SREN LM signal -> 3-D point cloud in JRC2018VNCF microns:
# 1. median3d denoise
# 2. keep top 50% of nonzero voxels by intensity
# 3. drop spatial outliers (largest connected component only)
# 4. decimate to 25k points sampled with prob proportional to
# intensity (bright voxels are likelier to be drawn)
# 5. alpha = (I/max)^2 with a 0.04 floor — dim voxels fade rather
# than blob into a flat green mass
ser <- nat::read.nrrd(nrrds[["sren"]])
ser <- mmand::medianFilter(ser, mmand::shapeKernel(c(3,3,3), type="box"))
storage.mode(ser) <- "integer"
cut <- as.integer(stats::quantile(ser[ser > 0], 0.50))
mask <- ser >= cut
cc <- mmand::components(mask, mmand::shapeKernel(c(3,3,3), type="box"))
mask <- !is.na(cc) & cc == which.max(tabulate(cc[!is.na(cc)]))
idx <- which(mask, arr.ind = TRUE)
ser_full <- data.frame(
X = (idx[, 1] - 1L) * 0.4 + 0.2,
Y = (idx[, 2] - 1L) * 0.4 + 0.2,
Z = (idx[, 3] - 1L) * 0.4 + 0.2,
I = as.integer(ser[mask]))
set.seed(42L)
N_VIZ <- 25000L
keep <- sample.int(nrow(ser_full), N_VIZ,
prob = ser_full$I / sum(ser_full$I))
# Forward-warp to BANC um (jrcvnc2018f um -> BANC nm -> BANC um).
ser_BANC <- Morpho::tps3d(
as.matrix(ser_full[keep, c("X","Y","Z")]),
bancr::jrcvnc2018f_to_banc_tpsreg$refmat,
bancr::jrcvnc2018f_to_banc_tpsreg$tarmat,
lambda = bancr::jrcvnc2018f_to_banc_tpsreg$lambda)
ser_pts <- data.frame(
X = ser_BANC[, 1] / 1000, Y = ser_BANC[, 2] / 1000,
Z = ser_BANC[, 3] / 1000, I = ser_full$I[keep])
# Depth-coded hue: green ramp (dark teal -> spring green -> lime).
# Single hue family so the cloud reads as the SREN signal, with depth
# layers separated by lightness. Combined with alpha by intensity
# (gamma 1.5) and back-to-front draw order.
ser_pal <- scales::col_numeric(
palette = c("#0a3d2a", "#147a4a", "#1f8b1f", "#7be37b", "#d8ff7f"),
domain = range(ser_pts$Z))
ser_pts$col <- ser_pal(ser_pts$Z)
hi <- max(ser_pts$I)
ser_pts$alpha <- pmax(0.05, (ser_pts$I / hi) ^ 1.5)
ser_pts <- ser_pts[order(-ser_pts$Z), ] # back-to-front
# (c) Tight zoom: union of all top-6 mesh bbox + SREN bbox (in X, Y, Z).
mesh_bb <- do.call(rbind, lapply(meshes_um, function(m) {
apply(nat::xyzmatrix(m), 2, range)
}))
xlim <- range(c(mesh_bb[, 1], ser_pts$X)) + c(-15, 15)
ylim <- range(c(mesh_bb[, 2], ser_pts$Y)) + c(-15, 15)
zlim <- range(c(mesh_bb[, 3], ser_pts$Z)) + c(-10, 10)
# 90 deg about X to view from the side; nat.ggplot::geom_neuron
# always plots rotated X vs rotated Y, so this puts Z on the y-axis.
rot_xz <- matrix(c(1,0,0, 0,0,1, 0,1,0), nrow = 3, byrow = TRUE)
abbr <- function(x) {
if (is.na(x)) return("—")
x <- sub("ventral_nerve_cord_neurosecretory_cell", "VNC neurosecretory",
x, fixed = TRUE)
x <- sub("abdomen_motor_neuron", "abd motor neuron", x, fixed = TRUE)
x
}
mk_panel <- function(i) {
rid <- top$root_888[i]
fmt <- function(x) ifelse(is.na(x), "—", x)
title <- paste(
sprintf("rank %d — score %.3f", i, top$score[i]),
rid,
abbr(top$cell_class[i]),
fmt(top$cell_type[i]),
sprintf("NT: %s", fmt(top$neurotransmitter_predicted[i])),
sep = "\n")
base_theme <- theme_void(base_size = 11) +
theme(plot.title.position = "plot",
plot.title = element_text(hjust = 0.5, size = 9.5,
lineheight = 1.05),
plot.margin = margin(4, 8, 4, 8),
legend.position = "none")
# Main: top-down YX (looking along Z).
yx <- ggplot() +
geom_point(data = ser_pts,
aes(x = X, y = Y, alpha = alpha, colour = col),
size = 0.45, stroke = 0, show.legend = FALSE) +
geom_neuron(meshes_um[[rid]], cols = "#aa44cc", alpha = 0.7) +
scale_alpha_identity() +
scale_color_identity() + scale_fill_identity() +
coord_fixed(xlim = xlim, ylim = ylim, expand = FALSE) +
scale_y_reverse() +
labs(title = title) + base_theme
# Side: XZ (looking along Y) -- shows the dorso-ventral spread.
xz <- ggplot() +
geom_point(data = ser_pts,
aes(x = X, y = Z, alpha = alpha, colour = col),
size = 0.45, stroke = 0, show.legend = FALSE) +
geom_neuron(meshes_um[[rid]], cols = "#aa44cc", alpha = 0.7,
rotation_matrix = rot_xz) +
scale_alpha_identity() +
scale_color_identity() + scale_fill_identity() +
coord_fixed(xlim = xlim, ylim = zlim, expand = FALSE) +
base_theme
patchwork::wrap_plots(yx, xz, ncol = 1, heights = c(3, 1.4))
}
p <- patchwork::wrap_plots(lapply(seq_len(nrow(top)), mk_panel), ncol = 3) +
patchwork::plot_annotation(
title = "Top-6 BANC efferent hits for the SREN query (BANC native um)",
subtitle = paste0("BANC EM hit (purple) + SREN LM source ",
"(hue = depth, alpha = intensity), in BANC native space"))
ggsave("banc_colormip_sren_top_hits.png", p, width = 13, height = 13, dpi = 130)What we recovered
The top-6 BANC efferents are anatomically what we would expect for SREN candidates:
| rank | cell_class | cell_type | nerve | neurotransmitter |
|---|---|---|---|---|
| 1 | abdomen_motor_neuron | MNANmX | abdominal_nerve_trunk | gaba |
| 2 | ventral_nerve_cord_neurosecretory_cell | EN00B010 | abdominal_nerve_trunk | octopamine |
| 3 | abdomen_motor_neuron | MNad48 | abdominal_nerve_trunk | octopamine |
| 4 | abdomen_motor_neuron | MNad17 | abdominal_nerve_trunk | serotonin |
| 5 | ventral_nerve_cord_neurosecretory_cell | abd_Dsx_OA | abdominal_nerve_trunk | gaba |
| 6 | abdomen_motor_neuron | CV1007 | abdominal_nerve_trunk | octopamine |
All six have arborisation in the abdominal ganglion and an axon
leaving via the abdominal nerve trunk — the route SREN
neurons take to reach the rectal papillae. With the tuned parameters,
colorMIP and NBLAST now agree well on the region and on three
of the top-6 cells (MNad48, MNad17,
CV1007); the remaining slots are filled by larger
neurosecretory cells whose abdominal arbour overlaps the SREN blob in
2-D MIP but whose 3-D mass extends elsewhere. NBLAST in Step 8 is the
discriminating tiebreaker.
The panels are rendered in BANC native µm rather than JRCVNC2018F because the bancr TPS landmarks cap at JRCVNC2018F Y ≈ 504 µm while the SREN signal extends to Y ≈ 537 µm — warping BANC meshes back into F-space at the posterior tip extrapolates ~30 µm short and produces misleadingly disjoint panels. Visualising in BANC native space (no extrapolation either way) shows the true 3-D relationship between query and hit. SREN points are coloured by depth (Z) — dark teal at one end of the slice through the abdominal ganglion, lime at the other — with alpha by intensity, so the cloud reads as 3-D layers rather than a flat mass.
Step 7 — Open each match in Spelunker
To inspect a match interactively we ship the SREN LM signal as a
Neuroglancer precomputed layer in BANC voxel space,
then build a per-match scene URL that turns on the BANC
segmentation proofreading layer with that match’s
root_id selected.
The precomputed layer at
gs://lee-lab_brain-and-nerve-cord-fly-connectome/light_level/kdrc/SREN_female_aligned240721_to_BANC.ng
was made by warping sren_female_sren.nrrd (JRCVNC2018F
microns) into BANC voxel space (2622 × 2950 × 789 @ 0.4 µm) with the
bancr Elastix VNC transform, then converting via
nrrd_to_precomputed().
# Re-create the BANC-space NRRD (one-off, ~5 min):
# Use bancr::vnc_240721 transformix params (BANC_to_template.txt)
# to bridge JRCVNC2018F -> BANC voxel grid, then cast to uint8.
# We then publish the precomputed layer:
nrrd_to_precomputed(
input = "sren_female_in_BANC_VNC.nrrd",
output = paste0("gs://lee-lab_brain-and-nerve-cord-fly-connectome/",
"light_level/kdrc/SREN_female_aligned240721_to_BANC.ng"),
resolution = c(400L, 400L, 400L),
data_type = "uint8",
encoding = "raw")
LM_GS <- paste0("precomputed://gs://lee-lab_brain-and-nerve-cord-fly-",
"connectome/light_level/kdrc/",
"SREN_female_aligned240721_to_BANC.ng")
BASE <- paste0("https://spelunker.cave-explorer.org/#!middleauth+",
"https://global.daf-apis.com/nglstate/api/v1/",
"6450802162925568")
build <- function(rid) {
bancr::banc_lm_scene(
lm_url = LM_GS,
layer_name = "KDRC SREN (female)",
range = c(1, 60),
opacity = 0.55,
blend = "additive",
ids = as.character(rid),
url = BASE,
shorten = TRUE)
}
top$ngl_url <- vapply(top$root_888, build, character(1))
head(top[, c("root_888", "score", "cell_type", "ngl_url")])Each URL opens the BANC scene with the matched cell selected and the SREN LM volume rendered as an additive overlay in green. The colorMIP top-6 links are pinned in a table at the bottom of this vignette.
Step 8 — Cross-check with NBLAST
Color-MIP search is fast and 2-D. NBLAST is 3-D, slower, and uses geometry directly — we can use it to cross-check the colorMIP ranking on the top-K candidates.
library(nat.nblast)
# (a) Re-score colorMIP top-100 with NBLAST. Fetch L2 dotprops in
# BANC microns for the candidate set.
candidates <- head(res$root_888, 100L)
banc_dps <- bancr::banc_read_l2dp(candidates)
# (b) Build the SREN dotprops *intensity-weighted*: sample 50k voxels
# from the largest connected component, with sampling probability
# proportional to voxel intensity. Bright voxels are over-represented
# in the resulting vector field, so the SREN dotprops is dominated by
# real signal rather than dim halo. This is the LM analogue of
# weighting an EM skeleton by node thickness.
N_NBLAST <- 50000L
keep <- sample.int(nrow(ser_full), N_NBLAST,
prob = ser_full$I / sum(ser_full$I))
ser_pts_F <- as.matrix(ser_full[keep, c("X","Y","Z")])
ser_BANC_nm <- Morpho::tps3d(
ser_pts_F,
bancr::jrcvnc2018f_to_banc_tpsreg$refmat,
bancr::jrcvnc2018f_to_banc_tpsreg$tarmat,
lambda = bancr::jrcvnc2018f_to_banc_tpsreg$lambda)
ser_dp_BANC <- nat::dotprops(ser_BANC_nm / 1000, k = 5L)
# (c) Forward + reverse NBLAST (fcwb scoring matrix), averaged.
fwd <- nat.nblast::nblast(ser_dp_BANC, nat::as.neuronlist(banc_dps),
smat = nat.nblast::smat.fcwb, normalised = TRUE)
rev <- nat.nblast::nblast(nat::as.neuronlist(banc_dps), ser_dp_BANC,
smat = nat.nblast::smat.fcwb, normalised = TRUE)
nb <- data.frame(root_888 = names(banc_dps),
nblast_mean = as.numeric((fwd + rev) / 2))
nb <- nb[order(-nb$nblast_mean), ]
head(nb, 6)In BANC native µm, NBLAST hits show clear 3-D overlap between the
SREN cloud and the EM mesh — the true match the colorMIP search was
approximating. Rank 1 is MNad48 / octopamine at score
0.461; a second MNad48 cell rises from cMIP rank 14 to
NBLAST rank 3 (intensity-weighted decisively prefers it).
abd_Dsx_OA, CV1007, and MNad17
round out the top 6 — abdominal motor / neurosecretory cells whose 3-D
dendrite envelope coincides with the SREN signal. Spearman correlation
between the (tuned) cMIP and NBLAST scores on the 200-candidate pool is
ρ = 0.901.
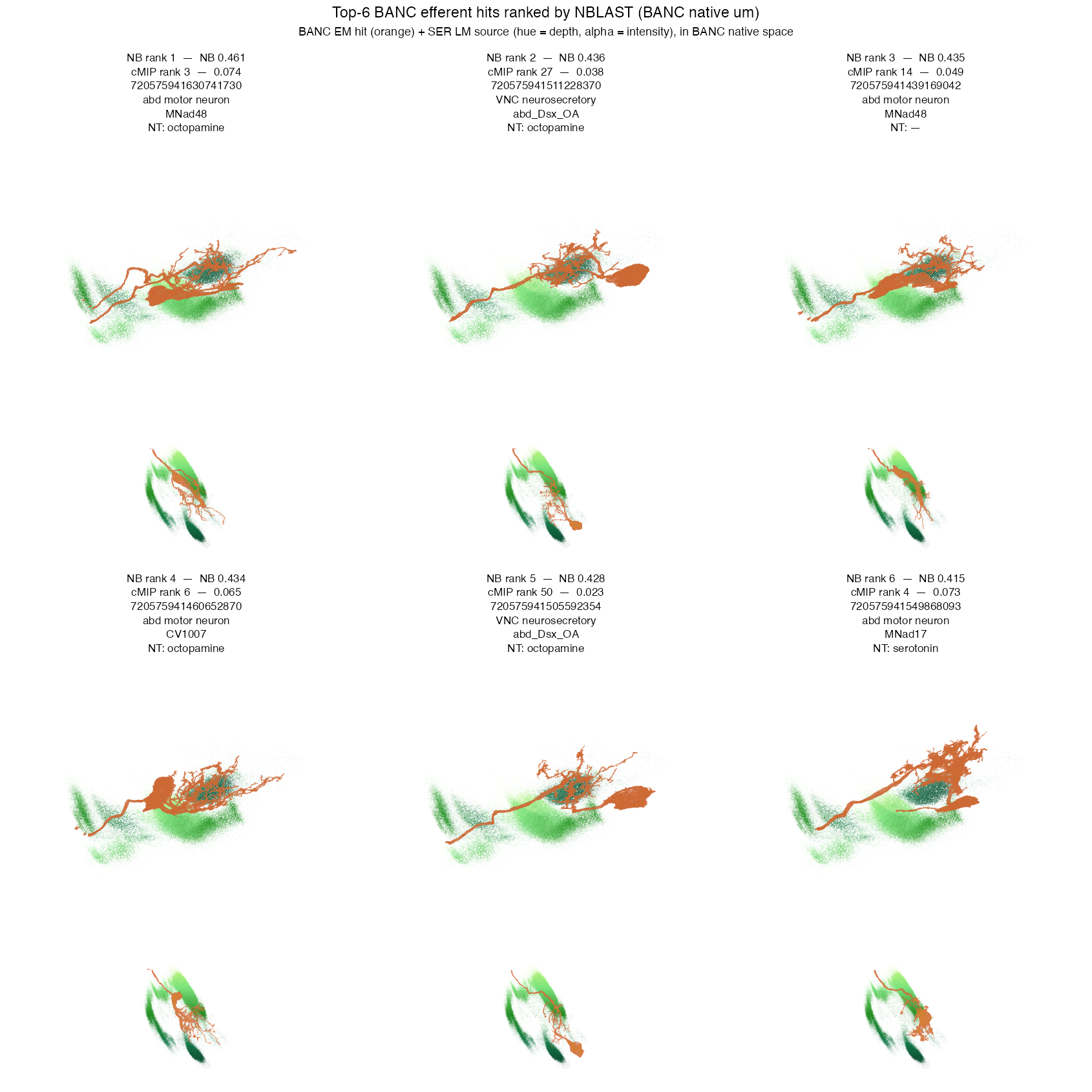
Consensus (averaged colorMIP + NBLAST rank, lower is better):
| consensus rank | root_id | cMIP | NBLAST | cell_type | NT |
|---|---|---|---|---|---|
| 1 | 720575941630741730 | 3 | 1 | MNad48 | octopamine |
| 2 | 720575941460652870 | 6 | 4 | CV1007 | octopamine |
| 3 | 720575941549868093 | 4 | 6 | MNad17 | serotonin |
The top consensus hit (MNad48, octopamine) is an
abdominal motor neuron projecting through the abdominal nerve trunk —
the expected neurotransmitter and routing for SREN cells supplying the
rectal papillae.
Pinned Spelunker URLs
For convenience, here are pre-built shortened Spelunker scenes for the top-6 colorMIP hits and the top-6 NBLAST hits. Each opens the BANC proofreading layer with the corresponding cell selected and the SREN LM volume rendered as a green additive overlay.
Top 6 by colorMIP (z_tolerance = 8, xy_shift = 3, mirror = FALSE)
| Rank | root_id | score | cell_type | NT | Spelunker |
|---|---|---|---|---|---|
| 1 | 720575941686473228 |
0.096 | MNANmX | gaba | open |
| 2 | 720575941628067385 |
0.091 | EN00B010 | octopamine | open |
| 3 | 720575941630741730 |
0.074 | MNad48 | octopamine | open |
| 4 | 720575941549868093 |
0.073 | MNad17 | serotonin | open |
| 5 | 720575941468851211 |
0.066 | abd_Dsx_OA | gaba | open |
| 6 | 720575941460652870 |
0.065 | CV1007 | octopamine | open |
Top 6 by NBLAST (intensity-weighted)
| Rank | root_id | score | cell_type | NT | Spelunker |
|---|---|---|---|---|---|
| 1 | 720575941630741730 |
0.461 | MNad48 | octopamine | open |
| 2 | 720575941511228370 |
0.436 | abd_Dsx_OA | octopamine | open |
| 3 | 720575941439169042 |
0.435 | MNad48 | — | open |
| 4 | 720575941460652870 |
0.434 | CV1007 | octopamine | open |
| 5 | 720575941505592354 |
0.428 | abd_Dsx_OA | octopamine | open |
| 6 | 720575941549868093 |
0.415 | MNad17 | serotonin | open |
Caveats
-
Threshold tuning. The R port and FIJI plugin use
semantically-different brightness thresholds (channel-sum vs
per-channel). Picking comparable settings is empirical; the validation
scatter in
inst/images/banc_colormip_r_vs_fiji.pngwas computed at Rthreshold = 60, FIJI1.threshold = 30 / 2.threshold = 30. -
Bridge accuracy. The Saalfeldlab JRC VNC H5 was
generated from population averages and is small in the body of the VNC
but can be imprecise at the periphery (where SREN axons leave). For
sub-voxel precision, register your specific NC82 channel onto
JRC2018VNCU_HR via Elastix multi-resolution affine + B-spline (the same
recipe as
lm_to_jrc2018u_elastixfor the brain). -
Library coverage. Only ~75 % of
flow == "efferent"BANC v888 cells (782 / 1035) had a colour-MIP rendered at the time of writing. As more get rendered, re-run Step 3 and the search. -
Mirror disabled for this query. The grid search
showed that
mirror = TRUEadds false-positive symmetric matches when the query is unilateral (as the SREN signal here is). For a bilateral query, setmirror = TRUEto score against the flipped library too.
Citations
- SREN identification (this query): Chae HS, Subay OH, Kim DH, Yoon SE, Kim YJ. Rectal Enteric Neurons Optimize Fecal Production in Response to Mating Signals. Conference talk, APDNC4, 2026. Department of Life Sciences, Gwangju Institute of Science and Technology (GIST), Gwangju 61005, Republic of Korea (corresponding author: Young-Joon Kim, kimyj@gist.ac.kr; Korea Drosophila Resource Center, GIST). Identifies a pair of sex-peptide-receptor neurons at the abdominal-ganglion tip whose efferents innervate the rectum and gate ROD production in response to mating; downstream targets ascend to the brain via Trans-Tango.
- BANC connectome: Phelps et al. (2024). Reconstruction of motor control circuits in adult Drosophila using automated transmission electron microscopy. Cell. https://doi.org/10.1016/j.cell.2020.12.013
- NeuronBridge / ColorMIP: Otsuna, Ito, Kawase. (2018). Color depth MIP mask search: a new tool to expedite Split-GAL4 creation. bioRxiv. https://doi.org/10.1101/318006
- JRC2018 VNC templates: Bogovic et al. (2020). An unbiased template of the Drosophila brain and ventral nerve cord. PLOS ONE. https://doi.org/10.1371/journal.pone.0236495
-
flybrainsPython package (Saalfeldlab JRC VNC H5 distribution): Schlegel et al., navis-org/navis-flybrains.