Design a split-GAL4 line for an EM neuron
Source:vignettes/design_split_gal4.Rmd
design_split_gal4.RmdGoal
Take a neuron from the hemibrain connectome, use NeuronBridge to find GAL4 and LexA lines that label it, and combine two of those into a split-GAL4 that targets just that one cell type. The example here uses LAL051, a local neuron in the LAL that drives descending motor circuits — a plausible target for sparse genetic access.
See the EM target in 3D
hemibrainr fetches the skeleton split into axon and
dendrite:
# Load a package we will need
library(hemibrainr) # to get this to work, please see:
# Find out neuron of interest
LAL051.info = neuprint_search("LAL051", field = "type")
LAL051.id = as.character(LAL051.info$bodyid)
LAL051.n = hemibrain_read_neurons(LAL051.id)
# Get the LAL
lal = neuprint_ROI_mesh("LAL(R)")
# Plot this neuron
nat::nopen3d()
plot3d(hemibrain.surf, col = "grey", alpha = 0.1)
plot3d(lal, col = hemibrain_bright_colors["red"], alpha = 0.1, add = TRUE)
hemibrain_view()
plot3d_split(LAL051.n, lwd = 2, soma = 400)
### Dendrite in blue, axon in orange, outouts in red, inputs in navy blue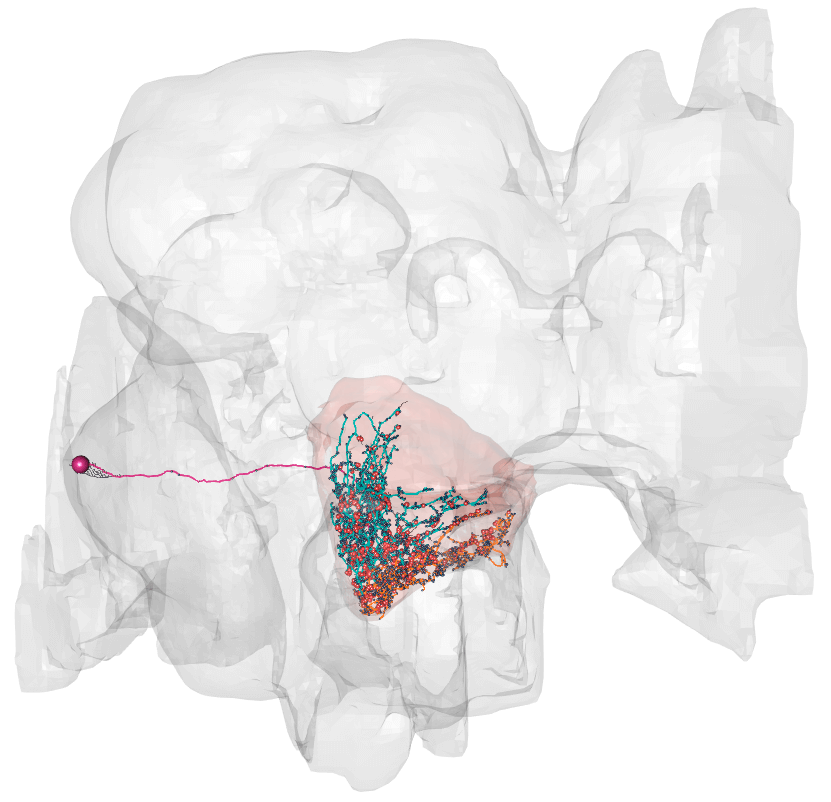
Render the target as a colour-depth MIP
NeuronBridge ranks LM lines by colour-depth MIP similarity to a 2-D EM render in which hue encodes z-depth (Otsuna et al. 2018). NeuronBridge already ships pre-computed MIPs for every hemibrain cell:
# What do we have on it?
nb.info = neuronbridge_info(LAL051.id)
View(nb.info)
# Let us now see the related MIP file
LAL051.mip = neuronbridge_mip(LAL051.id)
## This gets every MIP file associated with id
# Plot the MIP image in an rgl viewer
clear3d()
plot_mip(LAL051.mip)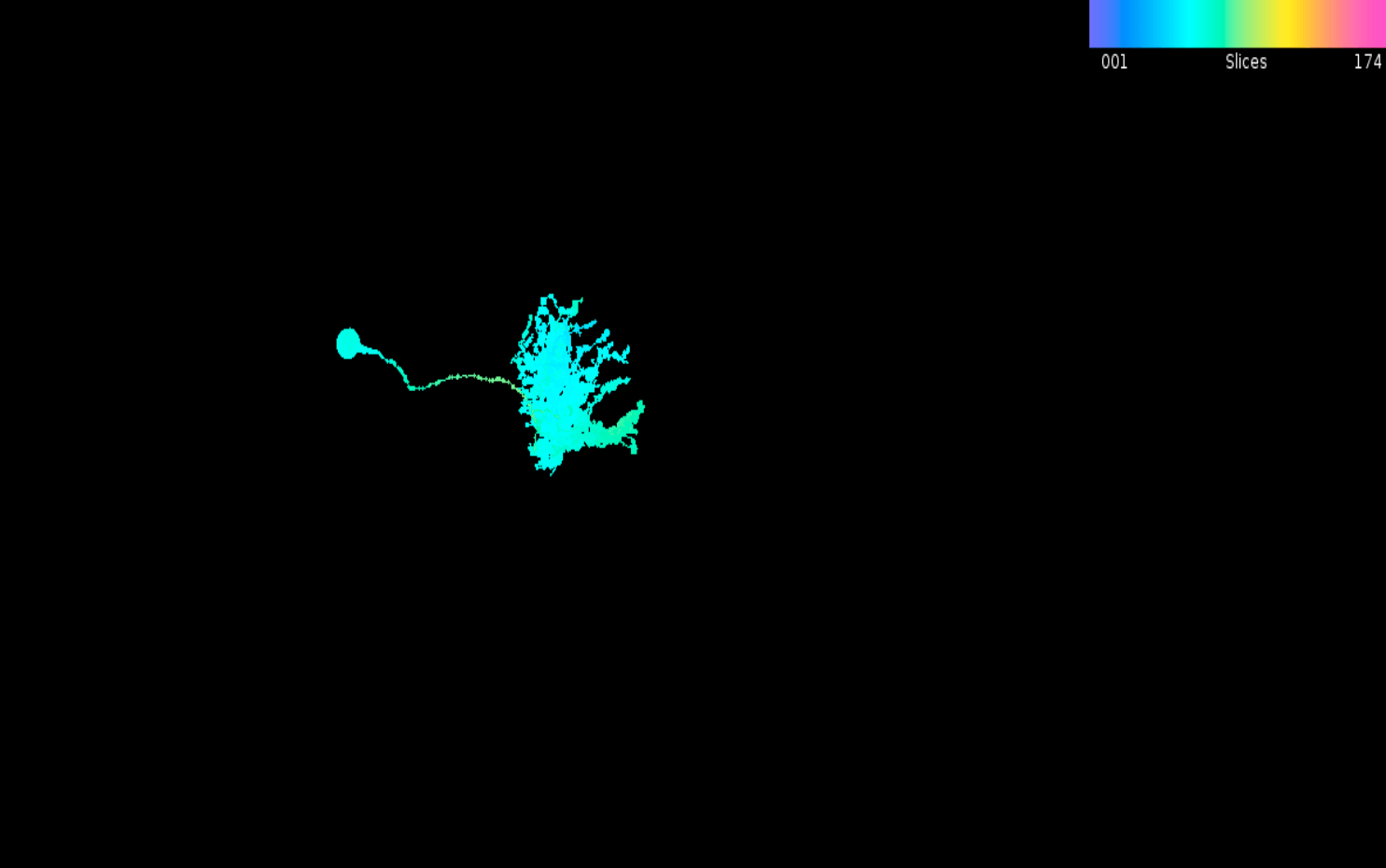
Search for lines that contain the target neuron
neuronbridge_search() returns the LM lines (mostly GAL4)
whose MIPs match the EM query, ranked by colour-depth score:
# Run search, looked at precomputed data
ns.s = neuronbridge_search(LAL051.id, threshold = 2000)
# scan hits
open3d()
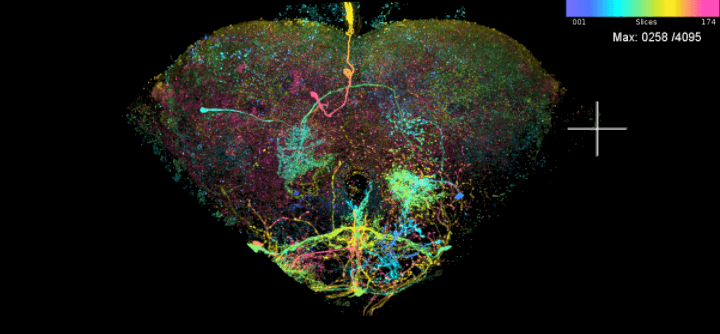
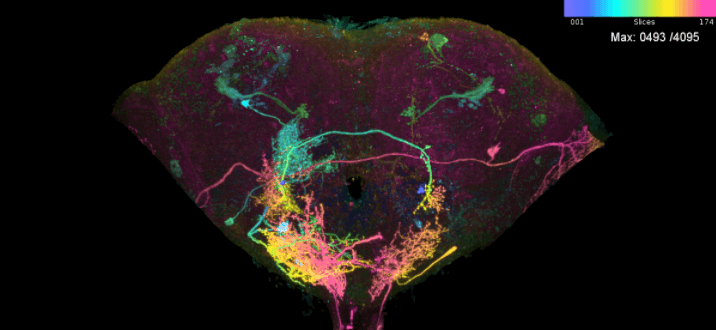
scan_mip(mips = ns.s, no.hits = nrow(ns.s))Top hits for LAL051: R58G03 and R71D08 —
two FlyLight enhancers that each include the cell, and that the
split-GAL4 design step below combines into a sparser line.
Predict the split
neuronbridge_predict_split() intersects the matching
neurons in two hemidrivers and reports the cells likely to survive in
their split-GAL4. Fewer surviving cells = sparser line.
line1 <- "R58G03"
line2 <- "R71D08"
potential.split <- neuronbridge_predict_split(
line1 = line1, line2 = line2,
threshold1 = 2000, threshold2 = 2000)
message("Predicted: ~", nrow(potential.split), " neurons in this split")
View(potential.split)